Research topics
Mobile genetic elements are more than a daily nuisance in the life of prokaryotes. While viruses reprogram and often kill cells in order to multiply, conjugative plasmids make cells addicted to plasmid-encoded anti-toxin molecules, preventing their disposal. Bacteria and Archaea defend themselves against invasion by these selfish invaders in a variety of ways including blocking the adsorption of viruses, preventing injection of their DNA, committing suicide upon infection and restriction-modification systems. One of the ways that microbes use to defend themselves is called CRISPR. In our lab we are interested in the molecular mechanism of defense systems in general, with a special focus on CRISPR.
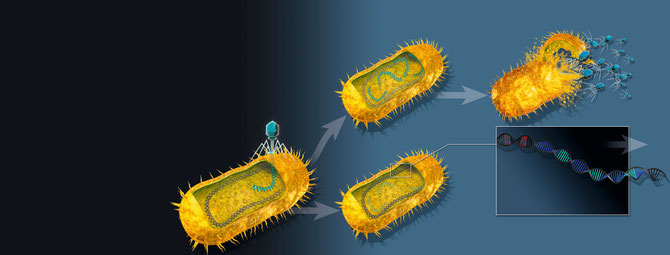
CRISPR-Cas
Since 2006 I have been interested in the mechanism of CRISPR immunity. The goal of my current lab is to understand how various antiviral defence systems such as CRISPR operate. This includes the formation of CRISPR memory, the assembly and function of CRISPR immune complexes, and the mode of DNA and RNA interference by CRISPR immune complexes and associated nucleases. In addition to these questions, we are interested in how the expression of these immune systems is regulated and controlled to withstand abundant bacteriophages.
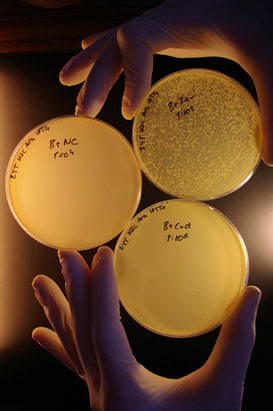
This photograph was taken in 2008 and shows the hands of Dr Magnus Lundgren holding plates showing for the first time that CRISPR immunity required specific guiding RNAs, Cascade and Cas3 to prevent viral infection. With randomly chosen guide RNAs (top right plate) the lambda virus can still infect and create plaques or clearing zones or plaques. This was a key experiment is determining the requirements of a CRISPR-Cas immune response.
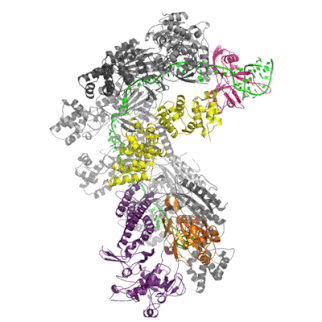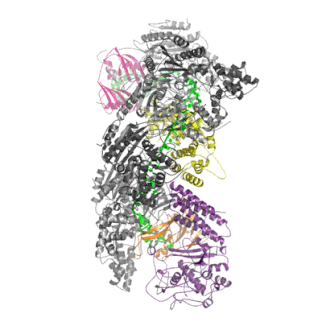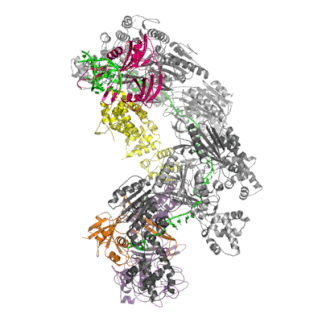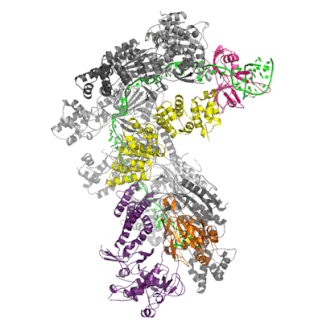
The Cascade complex is the key player in CRISPR immunity in Type I CRISPR-Cas immune systems. Discovered by my colleague Matthijs Jore in 2006 in Wageningen (Netherlands), the protein complex consists of 5 different Cas proteins and one crRNA (in green). The protein complex uses the crRNA as a guide to find and bind complementary DNA of the virus. The binding of Cascade to DNA flags it for for destruction by Cas3.

Once Cascade finds the target DNA, it recruits the nuclease Cas3, which will unwind and cleave the target DNA (in orange) before a virus can take control of the cell and replicate itself to many new copies.
The Cas1-Cas2 complex is involved in the formation of new CRISPR memories, by taking small fragments of viral DNA and integrating them between repeats in the host genome.

Bacteriophages
Viruses that infect Bacteria are called bacteriophages and they are arguably the most abundant and diverse biological entities on earth. We are interested in bacteriophage genome organisation, host range, and adaptations to host defence systems. We improve phages for phage therapy applications. Feel free to inquire about joining any of our bacteriophage related projects.
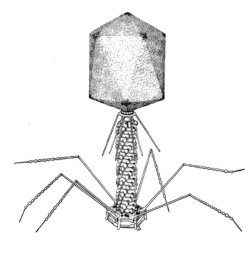
Bacteriophage T4 is one of the best studied bacteriophages to date. Within 22 minutes, T4 injects its DNA into the cell, degrades host DNA and RNA, replicates its own DNA and assembles into 100-150 phage particles. These are released into the environment by lysing the host cell from within. The phage stores its DNA under high pressure in the head of the phage particle, facilitating the ejection of the DNA into the cell.
Techniques and collaborations
We use a whole range of disciplines and techniques including Genetics, Biochemistry, Synthetic Biology, Next generation sequencing (MiSeq, MinION), Electron Microscopy, and High Throughput Screening and Selection schemes.
We collaborate with groups specialised in Metagenomics, Single Molecule spectroscopy, X-ray crystallography, and applications of Cas9 genome editing.
Further reading about CRISPR and Cas9
Host institute
Research interest
My lab is interested in the interaction between microbes and bacteriophages. We study the mechanisms that bacteria use to protect themselves from infections including CRISPR and other phage defense systems, and we explore the adaptations that viruses have evolved to avoid defence systems. We isolate and engineer bacteriophages for phage therapy applications.
Funding
Address
Kavli Institute of Nanoscience
Delft University of Technology
Applied Sciences Building (nr. 58) Office E1.480
Van der Maasweg 9
2629 HZ Delft
Netherlands






